_IN_JAK_STAT_PATHWAY_21.png)
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
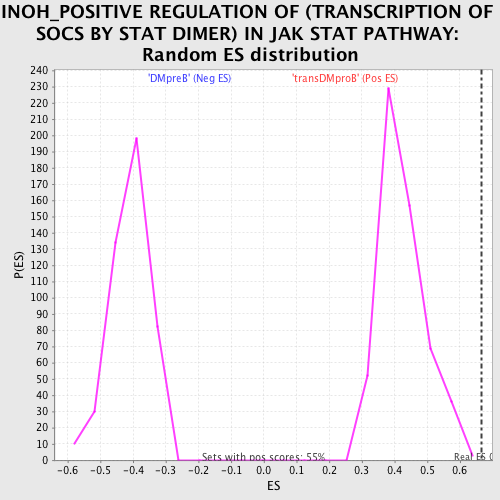
| GeneSet | INOH_POSITIVE REGULATION OF (TRANSCRIPTION OF SOCS BY STAT DIMER) IN JAK STAT PATHWAY |
| Enrichment Score (ES) | 0.66684705 |
| Normalized Enrichment Score (NES) | 1.5748032 |
| Nominal p-value | 0.0018315018 |
| FDR q-value | 0.13666041 |
| FWER p-Value | 0.59 |
_IN_JAK_STAT_PATHWAY_21.png)
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KIT | 16823 | 40 | 3.755 | 0.0671 | Yes | ||
| 2 | GFRA1 | 9015 | 56 | 3.423 | 0.1295 | Yes | ||
| 3 | STAT4 | 14251 9907 | 80 | 2.977 | 0.1831 | Yes | ||
| 4 | MAPK9 | 1233 20903 1383 | 89 | 2.817 | 0.2347 | Yes | ||
| 5 | MAPKAPK2 | 13838 | 165 | 2.313 | 0.2733 | Yes | ||
| 6 | MAP4K2 | 6457 | 210 | 2.155 | 0.3106 | Yes | ||
| 7 | JAK3 | 9198 4936 | 320 | 1.798 | 0.3379 | Yes | ||
| 8 | STAT3 | 5525 9906 | 396 | 1.605 | 0.3635 | Yes | ||
| 9 | CAMK2B | 20536 | 457 | 1.521 | 0.3883 | Yes | ||
| 10 | MAP3K3 | 20626 | 496 | 1.448 | 0.4129 | Yes | ||
| 11 | LCK | 15746 | 521 | 1.410 | 0.4377 | Yes | ||
| 12 | STAT1 | 3936 5524 | 531 | 1.404 | 0.4631 | Yes | ||
| 13 | MAP3K14 | 11998 | 645 | 1.197 | 0.4790 | Yes | ||
| 14 | MAP3K11 | 11163 | 734 | 1.072 | 0.4941 | Yes | ||
| 15 | MAPK14 | 23313 | 872 | 0.931 | 0.5038 | Yes | ||
| 16 | PDPK1 | 23097 | 914 | 0.896 | 0.5181 | Yes | ||
| 17 | PRKCH | 21246 | 1030 | 0.813 | 0.5269 | Yes | ||
| 18 | PRKCD | 21897 | 1031 | 0.813 | 0.5419 | Yes | ||
| 19 | BTK | 24061 | 1069 | 0.787 | 0.5544 | Yes | ||
| 20 | MAP3K5 | 20079 11166 | 1088 | 0.768 | 0.5676 | Yes | ||
| 21 | PAK4 | 17909 | 1147 | 0.717 | 0.5777 | Yes | ||
| 22 | MAP4K3 | 22887 | 1158 | 0.707 | 0.5902 | Yes | ||
| 23 | RIPK1 | 5381 | 1189 | 0.687 | 0.6013 | Yes | ||
| 24 | STAT5A | 20664 | 1207 | 0.676 | 0.6128 | Yes | ||
| 25 | MAP3K1 | 21348 | 1290 | 0.626 | 0.6199 | Yes | ||
| 26 | MKNK2 | 3299 9392 | 1390 | 0.578 | 0.6253 | Yes | ||
| 27 | JAK1 | 15827 | 1468 | 0.543 | 0.6311 | Yes | ||
| 28 | TEC | 16514 | 1558 | 0.496 | 0.6354 | Yes | ||
| 29 | STAT2 | 19840 | 1624 | 0.469 | 0.6406 | Yes | ||
| 30 | CAMK2G | 21905 | 1726 | 0.420 | 0.6429 | Yes | ||
| 31 | GSK3B | 22761 | 1874 | 0.364 | 0.6416 | Yes | ||
| 32 | AKT3 | 13739 982 | 1896 | 0.357 | 0.6471 | Yes | ||
| 33 | PTK2B | 21776 | 1904 | 0.355 | 0.6533 | Yes | ||
| 34 | MAP2K3 | 20856 | 1975 | 0.333 | 0.6556 | Yes | ||
| 35 | ACVR1 | 4334 | 1996 | 0.324 | 0.6605 | Yes | ||
| 36 | TYK2 | 12058 19215 | 2046 | 0.310 | 0.6636 | Yes | ||
| 37 | CDK5 | 16591 | 2088 | 0.297 | 0.6668 | Yes | ||
| 38 | CDK2 | 3438 3373 19592 3322 | 2297 | 0.243 | 0.6601 | Yes | ||
| 39 | CSK | 8805 | 2369 | 0.230 | 0.6605 | Yes | ||
| 40 | CDK4 | 3424 19859 | 2370 | 0.230 | 0.6647 | Yes | ||
| 41 | MAPK3 | 6458 11170 | 2440 | 0.214 | 0.6649 | Yes | ||
| 42 | IKBKE | 67 | 2498 | 0.200 | 0.6655 | Yes | ||
| 43 | MAP3K8 | 23495 | 2611 | 0.176 | 0.6627 | Yes | ||
| 44 | CDK6 | 16600 | 2683 | 0.159 | 0.6618 | Yes | ||
| 45 | IGF1R | 9157 | 2687 | 0.158 | 0.6646 | Yes | ||
| 46 | RPS6KA1 | 15725 | 2699 | 0.155 | 0.6668 | Yes | ||
| 47 | PAK1 | 9527 | 2858 | 0.127 | 0.6606 | No | ||
| 48 | MAP3K7 | 16255 | 3003 | 0.105 | 0.6548 | No | ||
| 49 | RAF1 | 17035 | 3038 | 0.101 | 0.6548 | No | ||
| 50 | ACVR2B | 19276 | 3128 | 0.092 | 0.6517 | No | ||
| 51 | MAP2K1 | 19082 | 3174 | 0.087 | 0.6509 | No | ||
| 52 | MAPK13 | 23314 | 3189 | 0.086 | 0.6517 | No | ||
| 53 | FGFR2 | 1917 4722 1119 | 3240 | 0.081 | 0.6505 | No | ||
| 54 | PDGFRB | 23539 | 3339 | 0.072 | 0.6465 | No | ||
| 55 | LTK | 14467 | 3451 | 0.063 | 0.6417 | No | ||
| 56 | CSNK1E | 6570 2211 11332 | 3467 | 0.062 | 0.6420 | No | ||
| 57 | ARAF | 24367 | 3602 | 0.052 | 0.6357 | No | ||
| 58 | CAMK2A | 2024 23541 1980 | 3717 | 0.046 | 0.6304 | No | ||
| 59 | ALK | 8574 | 3977 | 0.034 | 0.6170 | No | ||
| 60 | ACVR2A | 8542 | 4102 | 0.029 | 0.6108 | No | ||
| 61 | FLT1 | 3483 16287 | 4212 | 0.026 | 0.6054 | No | ||
| 62 | BMPR2 | 14241 4081 | 4228 | 0.026 | 0.6051 | No | ||
| 63 | MAPK7 | 1381 20414 | 4301 | 0.024 | 0.6016 | No | ||
| 64 | ZAP70 | 14271 4042 | 4669 | 0.016 | 0.5821 | No | ||
| 65 | ACVRL1 | 22354 | 5159 | 0.011 | 0.5558 | No | ||
| 66 | TGFBR1 | 16213 10165 5745 | 5364 | 0.009 | 0.5449 | No | ||
| 67 | PRKG1 | 5289 | 5810 | 0.007 | 0.5210 | No | ||
| 68 | MAP3K2 | 11165 | 6028 | 0.006 | 0.5093 | No | ||
| 69 | RIPK2 | 2528 15935 | 6498 | 0.004 | 0.4840 | No | ||
| 70 | BMPR1B | 8661 15152 4451 | 6558 | 0.004 | 0.4809 | No | ||
| 71 | PRKAR2B | 5288 2107 | 7813 | 0.000 | 0.4131 | No | ||
| 72 | FGFR1 | 3789 8968 | 8129 | -0.000 | 0.3961 | No | ||
| 73 | PRKCI | 9576 | 8313 | -0.001 | 0.3862 | No | ||
| 74 | RPS6KA2 | 9759 9758 | 8473 | -0.001 | 0.3776 | No | ||
| 75 | PAK3 | 9528 | 8538 | -0.001 | 0.3742 | No | ||
| 76 | ITK | 4934 | 8570 | -0.001 | 0.3725 | No | ||
| 77 | STAT5B | 20222 | 8729 | -0.002 | 0.3640 | No | ||
| 78 | NGFR | 5174 | 9182 | -0.003 | 0.3396 | No | ||
| 79 | MOS | 16254 | 9303 | -0.003 | 0.3332 | No | ||
| 80 | FYN | 3375 3395 20052 | 9586 | -0.004 | 0.3180 | No | ||
| 81 | MAPKAPK5 | 16387 | 9650 | -0.004 | 0.3146 | No | ||
| 82 | PRKCE | 9575 | 10001 | -0.005 | 0.2958 | No | ||
| 83 | KDR | 16509 | 10086 | -0.005 | 0.2913 | No | ||
| 84 | AKT2 | 4365 4366 | 10087 | -0.005 | 0.2914 | No | ||
| 85 | PRKCQ | 2873 2831 | 10097 | -0.005 | 0.2911 | No | ||
| 86 | IKBKG | 2570 2562 4908 | 10346 | -0.006 | 0.2777 | No | ||
| 87 | PLK1 | 9590 5266 | 10979 | -0.008 | 0.2437 | No | ||
| 88 | SRC | 5507 | 11341 | -0.009 | 0.2243 | No | ||
| 89 | BMPR1A | 8660 4450 | 11430 | -0.010 | 0.2198 | No | ||
| 90 | CDK7 | 21365 | 11644 | -0.011 | 0.2084 | No | ||
| 91 | PRKCA | 20174 | 11674 | -0.011 | 0.2071 | No | ||
| 92 | PRKAR1B | 16320 | 11799 | -0.012 | 0.2006 | No | ||
| 93 | JAK2 | 23893 9197 3706 | 11807 | -0.012 | 0.2004 | No | ||
| 94 | PDGFRA | 16824 | 11924 | -0.012 | 0.1944 | No | ||
| 95 | PRKACA | 18549 3844 | 11943 | -0.012 | 0.1936 | No | ||
| 96 | FGFR3 | 8969 3566 | 12054 | -0.013 | 0.1879 | No | ||
| 97 | MET | 17520 | 12124 | -0.013 | 0.1844 | No | ||
| 98 | EGFR | 1329 20944 | 12610 | -0.017 | 0.1585 | No | ||
| 99 | FGFR4 | 3226 | 12709 | -0.018 | 0.1535 | No | ||
| 100 | TGFBR2 | 2994 3001 10167 | 13012 | -0.021 | 0.1376 | No | ||
| 101 | RPS6KB2 | 7198 | 13037 | -0.021 | 0.1367 | No | ||
| 102 | FLT4 | 20904 1461 | 13136 | -0.022 | 0.1318 | No | ||
| 103 | AKT1 | 8568 | 13176 | -0.023 | 0.1301 | No | ||
| 104 | MAPK10 | 11169 | 13498 | -0.027 | 0.1132 | No | ||
| 105 | WEE1 | 18127 | 13773 | -0.032 | 0.0990 | No | ||
| 106 | INSR | 18950 | 13984 | -0.036 | 0.0883 | No | ||
| 107 | MAPK11 | 2264 9618 22163 | 14100 | -0.039 | 0.0828 | No | ||
| 108 | NLK | 5179 5178 | 14530 | -0.051 | 0.0606 | No | ||
| 109 | PRKG2 | 74 | 14843 | -0.066 | 0.0449 | No | ||
| 110 | RPS6KA3 | 8490 | 14937 | -0.071 | 0.0412 | No | ||
| 111 | MAP3K12 | 6454 11164 | 15194 | -0.089 | 0.0290 | No | ||
| 112 | PRKCZ | 5260 | 15276 | -0.095 | 0.0263 | No | ||
| 113 | MAP2K6 | 20614 1414 | 15369 | -0.106 | 0.0233 | No | ||
| 114 | SYK | 21636 | 15422 | -0.112 | 0.0226 | No | ||
| 115 | IRAK4 | 22379 11185 | 15529 | -0.124 | 0.0191 | No | ||
| 116 | MAP2K4 | 20405 | 15611 | -0.134 | 0.0172 | No | ||
| 117 | MKNK1 | 2504 | 15624 | -0.136 | 0.0191 | No | ||
| 118 | GRK6 | 6449 | 15684 | -0.144 | 0.0185 | No | ||
| 119 | MAP2K2 | 19933 | 15726 | -0.151 | 0.0191 | No | ||
| 120 | MNAT1 | 9396 2161 | 15837 | -0.169 | 0.0163 | No | ||
| 121 | PAK2 | 10310 5875 | 16141 | -0.223 | 0.0040 | No | ||
| 122 | MAPK8 | 6459 | 16226 | -0.245 | 0.0040 | No | ||
| 123 | IRAK1 | 4916 | 16274 | -0.256 | 0.0062 | No | ||
| 124 | IKBKB | 4907 | 16287 | -0.259 | 0.0103 | No | ||
| 125 | PLK3 | 15790 | 16505 | -0.311 | 0.0043 | No | ||
| 126 | RIPK3 | 21817 | 16528 | -0.316 | 0.0089 | No | ||
| 127 | CCNH | 7322 | 17056 | -0.491 | -0.0105 | No | ||
| 128 | MAPK12 | 2182 2215 22164 | 17159 | -0.534 | -0.0062 | No | ||
| 129 | CHUK | 23665 | 17214 | -0.556 | 0.0011 | No | ||
| 130 | PRKAR2A | 5287 | 17536 | -0.722 | -0.0029 | No | ||
| 131 | RYK | 19338 3870 | 17617 | -0.772 | 0.0070 | No | ||
| 132 | PTK2 | 22271 | 18532 | -2.547 | 0.0045 | No |
_IN_JAK_STAT_PATHWAY_22.png)